Marine ecosystems are being accelerated by human activities (such as climate change). Rising ocean temperatures lead to pathogen outbreaks, harmful algal blooms, and coral stress. These may cause severe consequences for marine ecosystems, human health, and aquaculture, forming a critical "One Health" issue. Monitoring key marine species provides valuable insights, but existing methods are resource-intensive, low-resolution, and unsuitable for frequent deployment. Harvard University researchers James J. Collins and Peter Q. Nguyen introduced a low-cost, field-deployable CRISPR biosensing platform for detecting DNA and RNA in marine organisms. Leveraging the programmability of CRISPR diagnostics in environmental biomonitoring, the platform demonstrates versatility across three climate-related indicators (Vibrio species, Pseudonitzschia species, and heat-stressed corals). A portable 3D-printed processor and incubator device enable direct processing of filter-captured samples with temperature control. Freeze-dried reagents, lateral flow assays, pipette operations, and a two-step multiplex detection workflow yield results within one hour without laboratory instruments, enhancing field testing convenience. Benchmarked with real pathogens and environmental seawater, the method confirms seawater tolerance and reliably detects 10⁸ colony-forming units of Vibrio pathogens per filter membrane, equivalent to 10² CFU per liter in filtered samples. This decentralized platform lowers the threshold for routine monitoring, enables early warnings for ecosystem disturbances, and supports the "One Health" initiative in marine science.
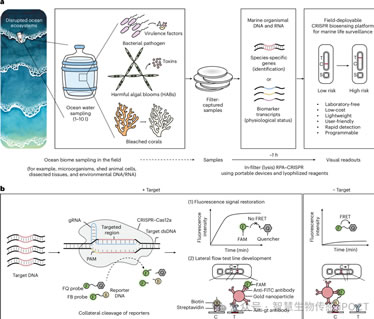
Figure 1: A field-deployable CRISPR biosensing platform for sustainable ocean monitoring. a. Climate change and anthropogenic pollution disrupt marine ecosystems, reshaping marine communities and leading to pathogen outbreaks, toxin-producing harmful algal blooms, and coral bleaching. CRISPR-based field-deployable biosensing technologies enable rapid, frequent, and user-driven ocean health monitoring. This can be achieved by detecting genetic markers of indicator species and assessing their physiological status through biomarker transcripts. These sensing methods are integrated into a lab-free workflow utilizing low-cost, lightweight 3D-printed devices capable of processing and detecting marine species in situ. b. Working mechanism of V-shaped CRISPR-Cas12a diagnostics. Cas12a targets double-stranded DNA under the guidance of complementary gRNA. Target recognition activates Cas12a's nonspecific nuclease activity, cleaving fluorescent/quencher-labeled probes (FQ probes, as shown in Figure 1) or FAM/biotin-labeled probes (FB probes, as shown in Figure 2), producing fluorescence or colorimetric LFA readouts, respectively.
Source: Sensor Expert Network



